RNA Translation: RNA makes Protein
In principle:
Translation of messenger
RNA (mRNA) takes place on ribosomes,
which include ribosomal RNA (rRNA),
with the help of transfer RNA (tRNA)
ribosomal RNA (rRNA)
rRNA + ribosomal protein  ribosomes
ribosomes
Structure of rRNA: stems
& loops
Structure of eukaryotic ribosome
subunits
Large Subunit + Small Subunit = 80S
monosome
A site (Aminoacyl), P site (Peptidyl), & E site (Exit) [APE or EPA complex]
transfer RNA
(tRNA)
adaptor molecule: ~30 tRNA types
2-dimensional 'cloverleaf' model
small: 75 ~ 90 nucs
stems & loops
D-loop & T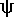 C-loop (
C-loop (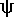 = pseudo-uridylic
acid)
= pseudo-uridylic
acid)
tRNA
characterized by 2o
modified bases
amino-acceptor stem
3' - ~~~~ CACCA - 3'
5' -~~~~ G
- 5'
anticodon loop
specificity of tRNA
for mRNA determined by 3-ribonucleotide sequence
3-dimensional
structure an "L"
D- & T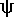 C-loops fold back on
each other
C-loops fold back on
each other
Charged tRNA: aminoacyl synthetase(x) forms ester linkage between
3'-A of
amino-acceptor stem of tRNA(x)
& COOH of amino acid(x)
~20
synthetase
types 'recognize' correct
anticodon loop
isoacceptance:
one-to-one correspondence between synthetase & amino
acid
RNA Translation: Protein
Synthesis
Ribosomes "read" mRNA & assemble polypeptide according to the Genetic Code
Initiation at start codon (AUG)
SSU binds
at Shine-Delgarno sequence (-6
nucs)
Multiple
complexes form on a single mRNA: polysome (polyribosome)
tRNAmet always added first [N-formyl-methionine in prokaryotes]

In
simplified form
Elongation: addition of amino acids according to Genetic
Code
P site amino acid transferred to A site amino acid
uncharged tRNA released from P site, passes
to E site
amino end
of initial met remains unchanged
and so
on ...
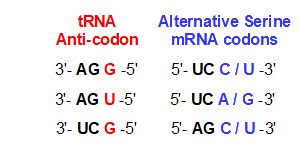
"Wobble": pairing of codon / anticodon
goes 5' 3'
on codon
3'
on codon
last position can miss-pair
with either purine / pyrimidine
Fewer tRNA species needed:
Ex.: three tRNAser species for six codons

Termination: release of polypeptide
stop codon (UAG, UAA, or UGA) enters A site
release factor
cleaves polypeptide from terminal tRNAn
polypeptide product: C
- lys - pro
- gly - phe
- met - N
Griffiths et
al. (1996) Fig.
13-7 is a nice schematic summary (HOMEWORK #11)

This is a logical,
not a biochemical, relationship:
Because mRNA is transcribed from the template strand,
it "looks like" the sense strand (except for
'U').
information
content of the DNA sense
strand and mRNA are identical
Protein sequences can be read directly from DNA:
Read the sense strand
in the 5' 3' direction,
3' direction,
Substitute 'T' for 'U' in the code table [or in
your head]
Computer programs ( MEGA, etc.)
do this automatically
There are three
reading frames on either strand
X
two 5' 3'
strands
3'
strands ![]() six possible ways to
read dsDNA
six possible ways to
read dsDNA
Open
Reading Frames suggest protein sequences
Deducing protein
sequences from "shotgun" DNA sequences:
a major research activity
Bioinformatics: extraction
of information from large macromolecular data
sets
These clues are useful:
Remember that all coding sequences:
are read only in the 5' 3' direction
3' direction
begin with a "start" (AUG)
codon
end with a "stop" (UAG, UAA, or
UGA) codon.
Ex.: a typical exam problem is to identify a polypeptide of six amino
acids from a dsDNA molecule
But: in real life research, any
large sequence of eukaryotic
DNA
may not have start and (or) stop codons for complete
protein,
[and
most AUG codons
are not 'start'
codons]
and may be include an intron with one or more 'stop' triplets .
Do not assume
that dsDNA molecules read
left to right, on top strand
Homework #12:
Practice DNA
"Translation" problems; there's an App for that: RandORF
All text material © 2024 by Steven M. Carr
