Protein Structure & Function
In principle:
Proteins are polymers of amino
acids
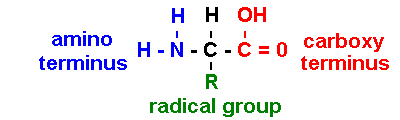
[sometimes NH3+ & COO- : depends on pH]
R = radical group determines biological properties: 20 types (note 1- & 3-letter codes)
|
Group
properties |
Three-
& Single-letter codes |
|
gly,
ala, val, leu, ile,
pro, met, phe, trp |
|
|
|
G
A V
L I
P M
F W |
|
ser, thr,
cys, tyr,
asn, gln |
|
|
|
S
T
C Y
N Q |
|
lys, arg,
his |
|
|
|
K
R
H |
|
asp, glu |
|
|
|
D E |
in vitro
dehydration of
carboxyl & amino termini forms peptide bond
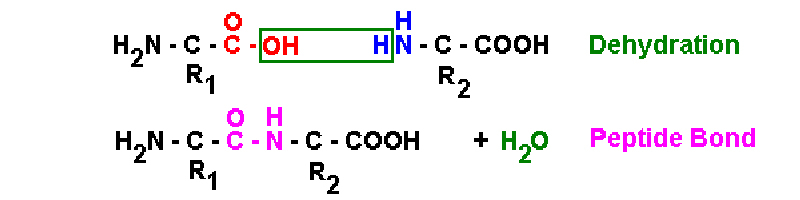
in vivo Peptidyl Transferase catalyzes
condensation reaction: H20
not lost
carboxyl
(C) terminus of growing
polypeptide in P site
cleaved from tRNA &
joined to amino (N) terminus of new amino
acid in A site
Repeating
backbone subunit [N - C(R) - C ] is amino acid residue
Primary Structure - order
of amino acid residues in polypeptide
20N possible orders with N residues
Secondary Structure - configuration of [-N-C(R)-C-]
backbone
alpha helix:
a
right-handed helix
beta-pleated-sheet: parallel / anti-parallel chains
both
stabilized by H-bonds
Tertiary Structure - 3-Dimensional folding
of backbone
Cys + Cys pairs form disulfide bridges
( -
S - S -)
Pro residues
form hydrophobic "corners"
hydrophilic residues
occur on exterior,
participate in reactions in aqueous environments
hydrophobic residues
occur in interior,
interact with membrane lipid bi-layer
Quaternary Structure - assembly of multiple subunits
dimers / tetramers / oligomers
e.g., hemoglobin is a tetramer:
two alpha + two beta chains
charged residues (Asp, Glu, Lys, Arg, His)
form ionic bonds bx
subunits
Post-translational
processing
Chemical modification of
amino acids
addition of formyl
group to Met ![]() fMet
fMet
Addition
of carbohydrate side
chains (glycosylation)
e.g., ABO blood group antigen proteins
Amino
acids may be cleaved out of primary structure
e.g., biologically active insulin is less than half
the primary
sequence
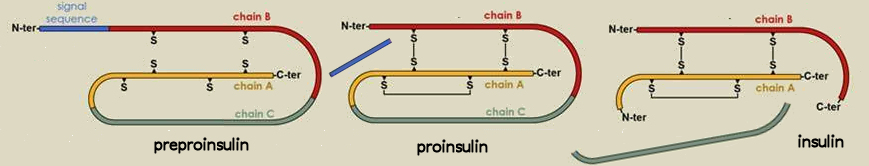
preproinsulin ![]() proinsulin
proinsulin ![]() insulin
insulin
(110 aa's)
(86 aa's)
(51 aa's)
Overview of protein function
Enzymatic catalysis of biological reactions
Substrates are bound in active sites: the Induced-Fit ModelLowered energy of activation
biological rxns occur at body temperature
with lower energy input
cf. Thermophillic hot spring bacteria live in 95oC H20
Identification of recurrent motifs allows
inferences about function
Helix - turn -
helix
motif binds Ca++
Zinc - finger motif binds major & minor
DNA grooves
Leucine Zipper motif
binds DNA and forms 'zippable' dimer
Other protein functions
Structural
Collagen constitutes 25% of human protein
Histones are the major components of chromosomes
Nucleic Acid binding
Polymerases, nucleases, helicases, ligases, etc.
Transport
Hemoglobin in
blood & myoglobin in muscle bind O2
Drosophila Genome Project
has cataloged 17,215
genes [Ensembl73
assembly]
~50% of Drosophila genes have human homologs
~75% of
human genetic disease-associated genes have Drosophila homologs
The
Human Genome
comprises 20,050 protein-coding genes: why so few?
Protein-coding exons may be
transcribed in different combination from different promoters
heterogeneous
nuclear RNAs (hnRNAs) may be spliced together (introns
spliced out) in different mRNA combinations
All text material ©2024 by Steven M. Carr