then
x=1 where L = life expectancy)
Short-term measures: "Life Table" parameters
rate of instantaneous increase (r) of a phenotype
recall
logistic
equation:
dN/dt = rN = rN (K-N)
/ K
where
K = carrying capacity
net reproductive rate: exp(r)
= er
r is "compound interest" on N
replacement rate (RO):
lifetime reproductive output
~
er(at low
density)
components of fitness:
traits
that contribute to survival
&
reproduction
Ex.: survivorship (expected survival time)
fecundity (# offspring at age x)
Adaptation
is
the
phenotypic
consequence
for populations of natural selection on
individuals
[cf.
adjust / acclimate]
Phenotypic
traits that change as a result of selection
are
sometimes
referred
to
as "adaptations"
or
"adaptive characters"
Life table analysis: survivorship and fecundity vary with age
lx
= prob. of survival from
birth to
age
x
(cumulative)
survivorship = probability
of
survival
to age x+1 from age x
mx
= fecundity (# offspring)
at age x
L
then ![]() (lx)(mx) exp(-rx) =
1
(in
a
stable
population,
(lx)(mx) exp(-rx) =
1
(in
a
stable
population,
x=1
where L = life expectancy)
L
Ro=![]() (lx)(mx)
(lx)(mx) ![]() replacement rate
replacement rate ![]() er at
low
density
er at
low
density
x=1
This
equation
is a discrete solution to the continuous
logistic
equation
Consider a population with two
demographic
phenotypes:
These
phenotypes
correspond to two reproductive 'strategies
iteroparous
strategy:
offspring produced over several seasons
semelparous
strategy:
offspring produced all in one season
A
survivorship
and fecundity schedule will compare their life histories
life table parameters can be measured experimentally
Under
'typical'
environmental
conditions,
survivorship
is 50% / year:
both
strategies
produce
2
young / female / lifetime
=>
both phenotypes are equally 'fit' [and N is
stable]
In 'good
times', survivorship increases to 75% / year:
iteroparous
strategy
produces
4
young / female / lifetime
semelparous
strategy
produces
3
young / female / lifetime
=>
iteroparous phenotype is 'more fit' [and N is
increasing]
In 'bad
times', survivorship decreases to 25% / year:
iteroparous
Ro = 0.72, semelparous Ro
= 1.00
=>
semelparous phenotype is 'more fit' [and N is
decreasing]
=> Population phenotypes will adapt to changing conditions
In a
favourable
environment, K increases:
e.g.,
productivity
of
meadow
increases
iteroparity
more
advantageous,
population density increases
In an
unfavourable
environment, r increases:
e.g.,
severity
of
winter
highly variable
semelparity
more
advantageous,
early reproduction favoured
K-strategy:
maintain population size N close to K
long-lived,
reproduce late, smaller # offspring, lots of parental care
E.g.,
many
bird
species,
primates (including Homo)
r-strategy:
maximize
growth
potential
r
short-lived,
reproduce early, larger # offspring, little parental care
E.g.,
most
invertebrates,
some
rodents
We can extend single-locus ![]() multilocus
multilocus ![]() quantitative models
quantitative models
p2:2pq:q2
W0,W1,W2
Mendel's
Laws
&
H-W
Theorem
![]()
![]()
![]()
normal
distribution fitness
function
high
heritability
Variation can be quantified
mean ![]() standard deviation:
standard deviation: ![]()
![]()
![]()
variance: ![]() 2
2
coefficient
of variation (CV)
=
(![]() /
/![]() )
x 100
)
x 100
CV
removes
size effect when comparing variance:
Ex.: Suppose X = whale
length
Y
= tail width
X = 100 ![]() 1.0
versus
Y = 1.0
1.0
versus
Y = 1.0 ![]() 0.1
0.1
CV of X =
1% CV
of
Y = 10%
Y is more variable, though ![]() X
is larger
X
is larger
Quantitative
variation follows "normal distribution"
(bell-curve) iff
Multiple
loci
are
involved
Each
locus
has
about
the same effect
Each
locus
acts
independently
[interaction variance (see below) is minimal]
Variation has two sources: genetic
(![]() G2)
&
environmental
(
G2)
&
environmental
(![]() E2)
variance
E2)
variance
phenotypic
variance ![]()
![]() P2
=
P2
= ![]() G2
+
G2
+ ![]() E2
+
E2
+ ![]() GxE2
GxE2
additive
variance ![]()
![]() A2
=
A2
= ![]() G2
+
G2
+ ![]() E2
E2 ![]()
heritability ![]() h2 =
h2 = ![]() G2/
G2/![]() A2
=
A2
= ![]() G2
/ (
G2
/ (![]() G2
+
G2
+ ![]() E2)
E2)
"heritability in the narrow sense": ignores ![]() GxE2
GxE2 ![]() interaction variance:
interaction variance:
Identical genotypes produce different phenotypes
in
different
environments.
Ex.: same breed of cows produces different milk yield on
different
feed
Artificial
breeding
indicates that
organismal variation is highly heritable
ex.:
Darwin's
pigeon breeding experiments
Artificial
selection
on agricultural species
Commercially
useful
traits
can
be improved by selective breeding
IQ scores in Homo:
h2![]() 0.7
0.7
[But:
IQ
scores
improve
with education: ![]() GxE2
is large]
GxE2
is large]
Offspring / Mid-parent
correlation
For
many
traits
in
many organisms:
CV = 5 ~ 10 %
h2 =
0.5 ~
0.9
[Read "Suggestions for using the Website" for used in this course]
Fitness
function
expresses relationship
between
genotype
& fitness
Function is a continuous
variable, rather than discrete values for W0,
W1,
& W2
=> Most traits vary & are
heritable.
Many traits
do
respond to 'artificial'
selection.
Many traits
should
respond to 'natural'
selection.
=> To demonstrate &
measure
Natural
Selection,
we must
show experimentally that heritable variation has consequences
for
fitness
<=
"Form
&
Function":
Organisms
typically
exhibit
engineering
criteria of "good
design"
Beak
variation in Hawai'ian honeycreepers (Drepaninae)
Beak
type
matches
food
type
Aerodynamics
of bat & bird flight
Slow
fluttering
bats
versus fast soaring birds
Wings
match
aerodynamic
principles
Assumption: Form & Function affect survival & reproduction
Persistence:
"Estimated time to extinction"
Are
long-lived
lineages
"better
adapted"?
Multituberculata
versus modern mammalian orders (3D
animation)
Order
persisted more than
twice
as long as any extant order
Ultimately
out-competed
by
Rodentia
Chondrichthyes
(sharks & rays) versus Teleosts
Body
form
is
unchanged
in 400 MY
Class
is
about
as
diverse now as at anytime in last 250 MY
Agnathan
orders [Hagfish & Lampreys] versus gnathostome
orders
Descendants
of
Ostracoderms,
500
MYBP (million
years
before
present)
Jawlessness
works
[ectoparasitism
is
probably secondary]
"Adaptive characters" cannot be separated from the organisms that bear them
Ex.:
We
typically
say "Hair & feathers evolved from scales".
But:
It
is
more
accurate to say:
"Reptiles (with scales) evolved into mammals (with hair)
and
birds
(with feathers)."
[and
this
isn't
completely
accurate either]
|
Agnaths (scaleless)
|
Ex.:
In
a
mammalogy
class, we might say
"The carnassial pair
evolved
from
the P4/M1
combination."
But:
it
is
more
accurate to say
"Carnivorous mammals evolved from insectivorous
ancestors.
The
carnassial
pair
is
adapted for slicing meat."
Quantitative
trait
distribution can be described as a bell curve
with
a
particular
mean
& variance:
What
happens
to
this
distribution under Selection?
(1) Directional Selection
Fitness
function has constant slope:
Trait
mean
shifted towards favored phenotype
trait
variance
unaffected
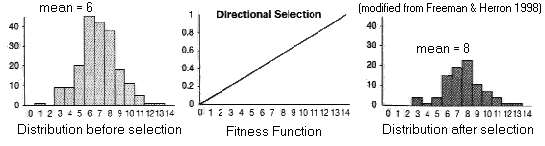
In
single-locus
models, the limit of selection is
Elimination
of
variation by fixation of favored allele
In quantitative
models,
rate is limited by
substitutional
genetic load:
"cost"
of
replacing
non-favored
allele (![]() "intensity" of selection)
"intensity" of selection)
"Soft"
selection
Mortality
is
density-dependent
In
'real'
populations: N(after)![]() N(before)
N(before)
Survivorship
is
proportional
to
fitness up to K: more realistic
Selection
will
affect
recruitment to next generation
Ex.: If the first-born dies of malaria, s/he will be
replaced.
More
births
occur
such
that N is continually "topped up".
Birth
of
succeeding
offspring
will maintain N near K
artificial selection on agricultural species
Gecko
lizard
(Aristelliger) has "suction pad" feet:
lamellar scale counts
increase
with
age
Darwin's
Finch
(Geospiza fortis) adapts to drought:
larger
birds
survive
because
of changes in seed
size
& hardness
(recall
that
size
is
heritable)
Developmental
canalization
limits extent of directional selection
Systems
are
controlled
by
multiple epistatic loci:
it
is
difficult
to
select on all loci simultaneously
Organisms
have
mechanical limits:
size
cannot
increase
indefinitely
Johanssen's bean experiment
Skull volume versus
birth
canal diameter in Homo
Phenotypes
are
not
infinitely
plastic:
[But: Eozostrodon
lineage
evolved
into whales & bats]
(2) Stabilizing
Selection (AKA truncation selection)
Fitness
function
has a "peak"
Trait
variance reduced around (existing) optimal phenotype,
trait
mean
unaffected
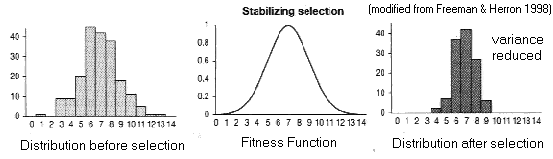
Limits:
elimination of variant alleles
or,
'weeding
out'
of
disadvantageous variants
homozygosity
at
multiple
loci:
difficult
iff variance due to recessive alleles (Lab
#1)
inbreeding depression: loss
of
'health'
in inbred lines
Examples:
Lab
#1: Elimination
of
non-cryptic
pepper moths (Biston)
Dark variants are
eliminated
rapidly in light environments
Light variants are
reduced
(more slowly) in dark environments (why?)
[This
may
look
like
an example of directional selection: why isn't it?]
Cold
shock in house sparrows (Passer) (Bumpus 1898)
Animals
that
die
are
at extremes of distribution
Birth weight
in Homo (Karn & Penrose 1951)
Modal
birth weight
is
optimum for survival
(3) Diversifying
Selection (two kinds)
There
is a lot of variation: does selection explain it?
(A) Balancing
Selection:
Fitness
function has more than one peak (multi-modal)
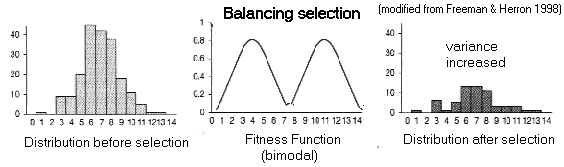
Trait
variance
increases
polymorphic["strict
sense"]: variation maintained within populations
Ex.:
cornsnakes, tomatoes,
bell
peppers, snails,
scallops
polytypic:
variation distributed among populations
Ex.:
shell patterns in Cepaea
snails
fraction
of
dark
/
banded shells varies with substrate
Limits:
segregational
genetic load:
loss
of reproductive potential due to production of less fit
homozygotes
In
Lab #1, Exercise #2, about
1/3 of
population "dies" in
malarial environment
Overdominance:
heterozygotes
have superior fitness at a locus
because
different
alleles
are
favoured in different environments
Examples:
sickle-cell
hemoglobin in Homo ('Contradictory' selection)
Leucine
Aminopeptidase
(LAP) & salinity
tolerance
in Mytilus mussels
hetero-dimers:
multimeric
enzymes
with
polypeptides
from different alleles
often
show
wider
substrate
specificity, kinetic properties (Vmax
& KM)
myoglobin in diving mammals
Heterosis:
heterozygosity at multiple loci improves general fitness
Hybrid
vigour: crossbreeding
of inbred
lines
improves fitness in F1
Marginal
epistasis: high 'Hobs' is 'good
for you'
Ex.:
correlation
between
phenotype
& genotype: antler
points in Odocoileus deer
Ex.:
fluctuating
asymmetry:
Acionyx
cheetahs
are lopsided
Maintaining polymorphic phenotypic variation by selection
Alternative
phenotypes favored in different environments
crypsis:
Cepaea
land
snails match background (Fig.
13-06)
Batesian
mimicry:
'Tasty' mimics converge on 'distasteful'
models
Viceroy
butterflies
(Limenitis) converge
on
Monarch (Papilio) butterflies
Mullerian
mimicry:
Distasteful
models
converge
on
each other,
different combinations evolve in different parts
of range
Heliconius
butterflies
(Futuyma 1997)
aposematic (warning)
colouration warns off predators (Mertensian mimicry]
Ex.:
scarlet
kingsnake
(nonvenomous)
mimics
coral
snake
(highly
venomous)
[black /
red / yellow pattern]
Frequency-dependent
selection:
Fitness
value
of
phenotype
varies with frequency
apostatic predation:
thrush
predation on Cepaea
'search
image'
changes
when
prey type becomes rare
'rare
male' effect: females prefer "different" male
Male
zebra
finches
with
artificial crest get more copulations (Fig.
20-13)
Sexual
Selection (Darwin 1871):
'exaggerated'
phenotypes
are
disadvantageous
somatically
but
are
favoured
in
competition for mates
secondary
sex
characteristics:
Sexual dimorphism in mallards,
peafowl,
& lions
Antlers in Cervidae are used in
male-male
combat
Tail displays in peacocks
attract
mates
'Runaway
sexual selection': the Madonna
/ Ozzy Osborne Effect
Females
choose
males
on
basis of some distinctive trait
Offspring
have
exaggerated
trait (males) & preference for
trait (females)
 selection
reinforces
trait & preference for trait simultaneously
selection
reinforces
trait & preference for trait simultaneously
New
phenotype
spreads rapidly in population
(B) Disruptive
selection
Fitness
function is a valley
Trait
variance
increases
(like
balancing), BUT polymorphism is
unstable
[Try NatSel with: q = 0.5, N = 9999, W0 = 1.0, W1 = 0.7, W2 = 1.0]
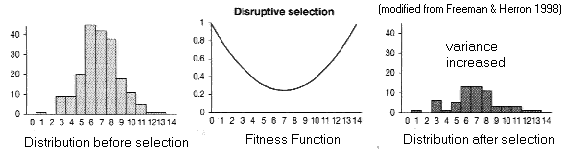
Polymorphism
can usually be maintained only temporarily:
One
of
the
phenotypes
will outcompete the other
unless
different phenotypes choose different niches (Ludwig Effect)
[and
then this becomes Balancing Selection]
Scutellar
bristles
in
Drosophila (Thoday & Gibson 1962)
Selection
for
'high #' versus
'low #' lines
=>
'pseudo-populations'
with
reduced interfertility
Might
disruptive
selection
contribute
to speciation?
Natural selection is ordinarily
defined
as
differential
survival
& reproduction of individuals:
Can
selection
operate on other biological units?
Can
such
selection 'oppose' individual selection?
Genic
(Gametic)
Selection
![]() Differential
survival
&
'reproduction' of alleles
Differential
survival
&
'reproduction' of alleles
Meiotic
Drive:
t-alleles in Mus
tt is
sterile
(W = 0)
Tt is
'tail-less'
(cf. Manx cats) (W
< 1)
t alleles are preferentially segregated into
gametes
(80~90%)
=>
f(t) is high in natural populations (40~70%)
even
though
it
is
deleterious to individuals
Kin
(Interdemic)
Selection
![]() Differential
survival
&
reproduction of related (kin) groups
(families)
Differential
survival
&
reproduction of related (kin) groups
(families)
Related
individuals
share
alleles:r = coefficient
of relationship [see
derivation]
offspring
&
parents
are
related by r = 0.50 [They
share
half their alleles]
full-sibs
"
"
r = 0.50
half-sibs
"
"
r = 0.25
first-cousins
"
"
r = 0.125
Inclusive
fitness (Wi)
of
phenotype
for
individual i
=
direct fitness of i + indirect fitness
of
relatives
j,k,l,...
Wi
= ai + ![]() (rij)(bij)
summed over all relatives j,k,l,...
(rij)(bij)
summed over all relatives j,k,l,...
where:
ai = fitness of i due
to
own phenotype
bij = fitness of j due to i's
phenotype
rij = coefficient of
relationship
of i & j
If
i & j are unrelated
warn:
Windividual
= 0.0 + (0.0)(1.0) = 0.0
don't
warn:
Windividual = 1.0 + (0.0)(0.0) = 1.0
 Such
behaviors
should not evolve among unrelated individuals
Such
behaviors
should not evolve among unrelated individuals
What
is
the
fitness
value in a kin group?
Wbrothers = 0.0
+
[(0.5)(1.0)
+ (0.5)(1.0)] = 1.0
Wcousins = 0.0 +
[8][(0.125)(1.0)]
= 1.0
 Such
behaviors
can evolve among
related individuals in (extended) family groups
Such
behaviors
can evolve among
related individuals in (extended) family groups
J.B.S.
Haldane (1892-1964):
"I would lay down my life for two
brothers
or eight cousins."
Parenting behaviour:
'Broken wing' display in
mother
birds
Mother sacrifices herself for (at
least two) offspring
Altruistic behaviour
(![]() "unselfish concern
for
others")
"unselfish concern
for
others")
'Alarm calls' in Belding ground
squirrels
(Spermophilus)
females
warn
more
in
related groups
Can
behaviours
to
help
unrelated individuals evolve?
Eusocial
insects
(Hymenoptera, Isoptera)
Haplodiploidy: females
diploid,
male
drones haploid
Females
workers
are
sterile
(Wi = 0): what is the selective
advantage?
related
to
queen
or
offspring by 1/2
related
to
sisters
by
3/4
 Care
for
sisters, don't have offspring
Care
for
sisters, don't have offspring
Natural Selection may be the most
misunderstood
concept in biology.
It is ...
(1) Not "Survival
of the Fittest"
Herbert
Spencer (1820 - 1903) "Social Darwinism"
the
"naturalistic fallacy": 'is'
=
'ought'
[Darwinian
theory
was
accepted
in part because
it
could
be
read
to support British imperial ambition]
not
phenotype-specific mortality
not predation (nor
inter-species
competition,
usually)
not
"Nature red
in tooth
and claw"
Darwin:
plants in desert 'struggle' for water
not
equivalent to population growth:
population
declined
in
semelparous
example
(2) Not equivalent to evolution
Natural
Selection may conserve existing types (stabilizing
selection).
Evolutionary
change ultimately requires new variation (mutation).
Migration,
population structure, genetic drift are important.
(3) Not a tautology (a self-evident statement; a circular argument)
"Why
do
they
survive?
Cuz
they're fit.
How
do
you
know
they're fit? Cuz they survive..."etc.
More
like
a syllogism (an if /
then
statement;
a logical consequence):
(![]() 2 &
2 & ![]() W
& h2) =>
W
& h2) => ![]() q
q
[cf. physics: F = M A depending on
how
Force,
Mass, & Acceleration are defined
arithmetic:
1
+
2
= 3 because I and II make III]
(4) Not "Mother Nature"
not
a force, not a thing that acts
[We
don't
say,
"Arithmetic causes one plus two to equal three."
We
might
say,
"One plus two equals three. That's arithmetic.]
not
good or bad (amoral)
no
noun
/ verb / object distinctions
[In
most
languages,
"nouns verb objects"
i.e., objects perform actions on other objects. Not.]
(5) Not teleological (goal-directed):
Evolution
does not have "goal", "direction", or "purpose"
(Homo sapiens are not the endpoint of evolution!)
Avoid
such phrases as "Natural Selection acts ..."
"in
order
to
...",
"for
the
purpose
of
...",
"so
that
...",
"because
its
trying
to
..."
Text material © 2012 by Steven M. Carr