Detection of Single-Nucleotide Polymorphisms (SNPs)

Detection of Single-Nucleotide Polymorphisms (SNPs)
Differences
between the DNA sequences of individuals and species are
due to alterations of the four-letter ACGT code. Almost all observed
differences are not newly-arisen "mutations" between the
previous and present generations, but rather persistent
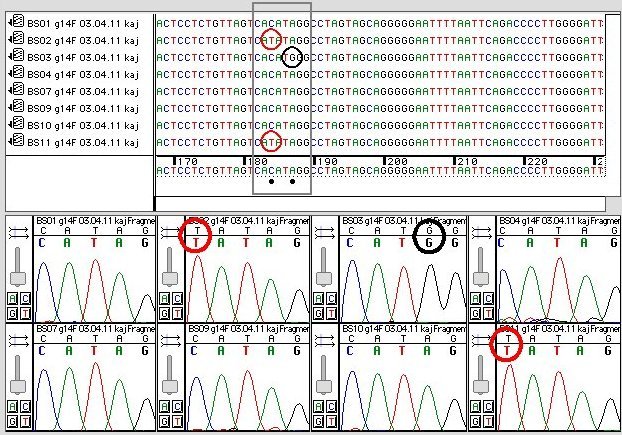
variations already present in populations. These are called single-nucleotide polymorphisms or SNPs. The image is a
portion of an Applied Biosystems Automated DNA sequencer
chromatogram.The top window
shows an 80-nucleotide stretch of DNA from a group of
eight Atlantic Cod. Black dots in the upper window flag
positions with at least one SNP variant. The grey
bracket frames an eight-base region with two SNPs, for
which the chromatogram data are shown in the lower
window. Most individuals have the sequence CACATAGG:
individuals BS02 & BS11
share a C T SNP variant at the third
position (CATATAGG),
and individual BS03 has a unique A
T SNP variant at the third
position (CATATAGG),
and individual BS03 has a unique A G SNP at the sixth position (CACATGGG).
Identification of such genetic differences among individuals or
species permits inferences about relationships among individuals
in time (genealogy) and space (phylogeography),
and the evolutionary history (phylogeny) of species. For
example, reconstruction of the pattern of SNP variation
shows that individuals BS02
and BS03 are
less closely related to each other than either is to individual
BS01.
G SNP at the sixth position (CACATGGG).
Identification of such genetic differences among individuals or
species permits inferences about relationships among individuals
in time (genealogy) and space (phylogeography),
and the evolutionary history (phylogeny) of species. For
example, reconstruction of the pattern of SNP variation
shows that individuals BS02
and BS03 are
less closely related to each other than either is to individual
BS01.