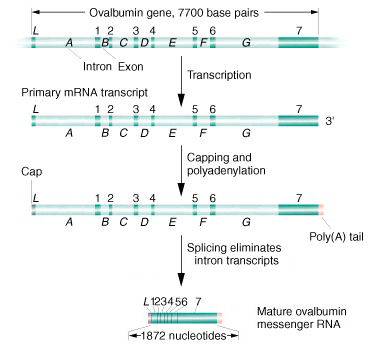
Post-Transcriptional processing of hnRNA
The primary hnRNA product
of chicken ovalbumin (ca. 7,700 b) is capped
and tailed.
The product is then spliced to join the segments of seven
exon transcripts
(1 - 7) into the mature mRNA
(1872 bp).
An important note on
terminology: Splicing of Exons &
Introns
'Splicing' is by definition the joining
of two things, for example the ends of two ropes. It is not
'splitting',
which is separating two things. However, in
molecular biology, ''splicing out' is often used
to refer to the removal of some hnRNA segments as a
consequence of splicing together the remaining
segments as mRNA.
'Exon' and
'intron' by definition refer
to regions of the DNA
that are respectively "expressed" and "intervening."
However, these terms are sometimes used loosely to refer to
the corresponding sequences in hnRNA that
are retained or removed, respectively, from
the final mRNA product.
These are more correctly called intron & exon transcripts,
as in the above diagram. The consequence is that the 5'![]() 3' sequences
of the DNA exons
in the sense
strand are the same as the corresponding mRNA exon transcripts, except for
substitution of U for
T. Thus the
corresponding amino acid sequences can be either 'read' directly from the DNA
sense strand, or 'translated' from the mRNA.
3' sequences
of the DNA exons
in the sense
strand are the same as the corresponding mRNA exon transcripts, except for
substitution of U for
T. Thus the
corresponding amino acid sequences can be either 'read' directly from the DNA
sense strand, or 'translated' from the mRNA.
Never use
"splicing to mean
splitting of the hnRNA transcript. Avoid thinking of exons as
the translated portions of the mRNA, while
recognizing that the word is often used that way, incorrectly.
In the figure above,
the phrase "Splicing eliminates intron transcripts"
correctly means that "Splicing, by joining together
the exon transcripts, results in the
elimination of intron transcripts." It does not
mean that splicing is the direct elimination of
intron transcripts.
Figure ©2000 by
Griffiths et al. ; text
©2026 by Steven M. Carr