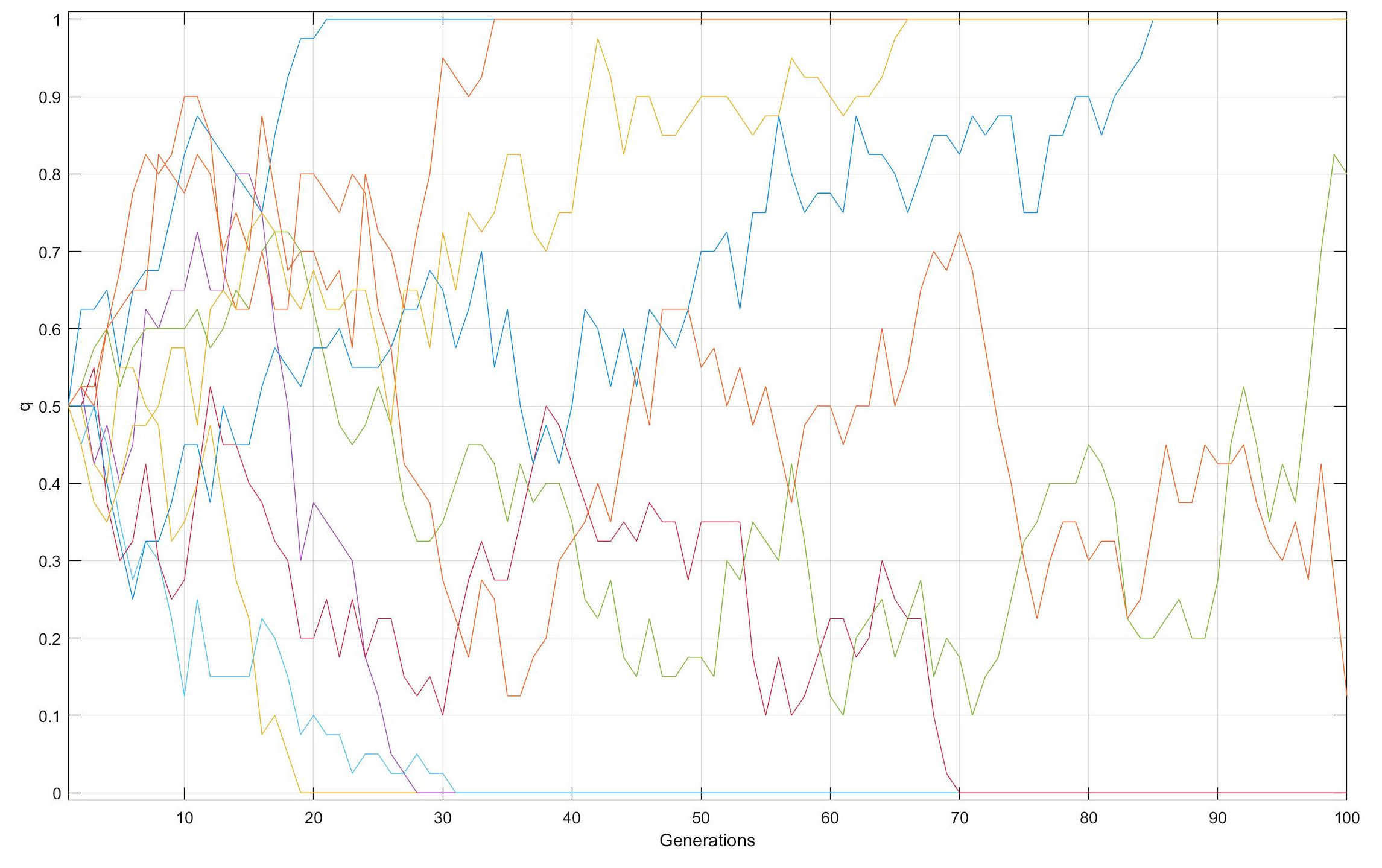
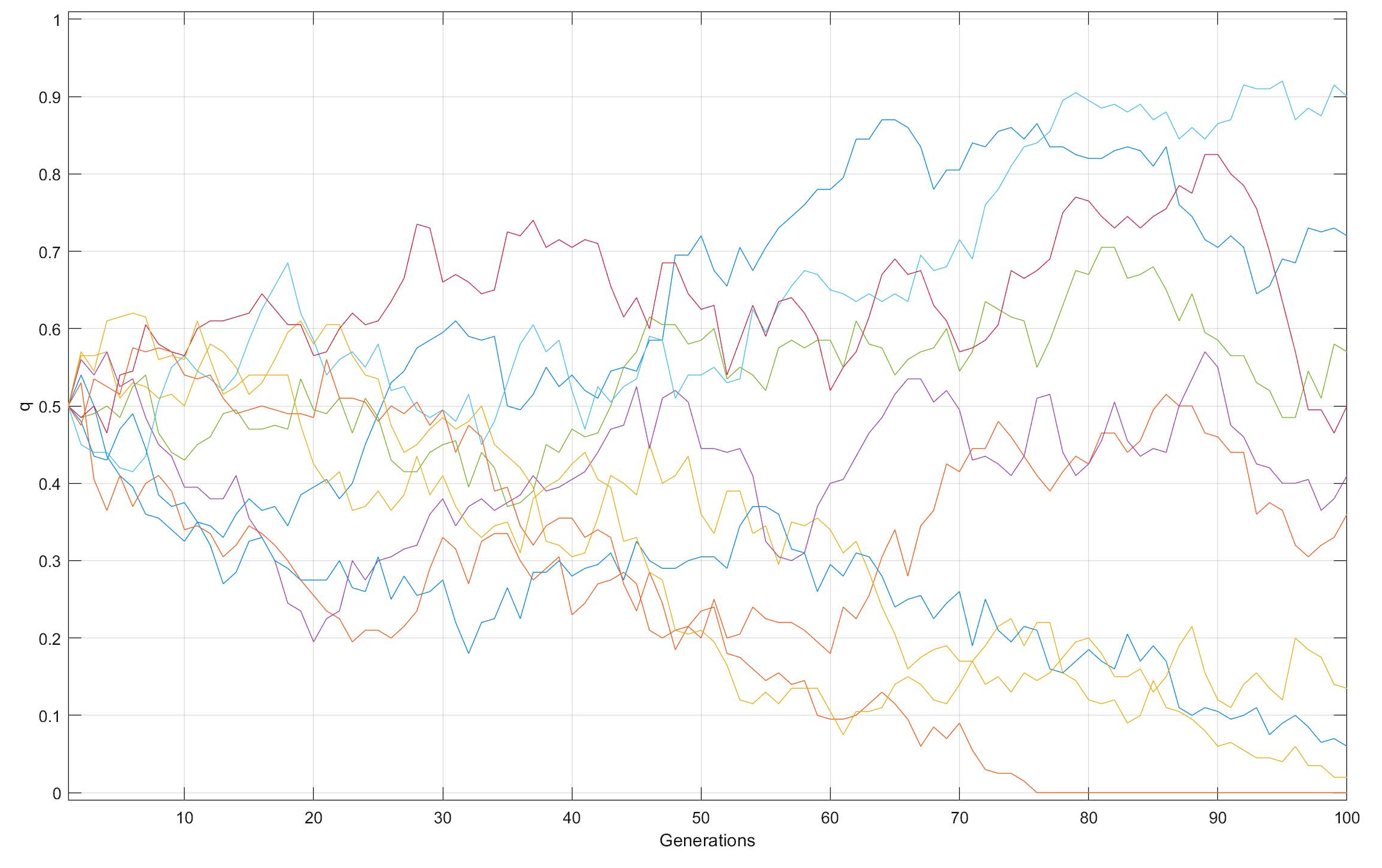
Genetic Drift over time in Smaller versus Larger populations
From an initial q = f(B) = 0.50 in each of 10 populations
with N = 20 [top] or 100 [bottom]
individuals, f(B) diverges at random over time from
the starting value. The divergence in more rapid among smaller
populations than larger. With N = 20, eight
populations have gone to either loss (q = 0.0)
(n = 4) or fixation (q = 1.0) (n = 4)
after 100 generations, whereas with N = 100 after 100
generations, all of the populations save one retain both
alleles, in the range of q = 0.01 ~ 0.90.
Allele frequencies continue to vary at random so long as 0 < q < 1: q = 0.0 or 1.0 are absorbing barriers, which once reached end further change. Fate of q in a population is not predictable: between g = 70 & 100 with N = 100, dark orange q goes from 0.70 to 0.15, and light green q from 0.10 to 0.80
HOMEWORK: Repeat the simulations with the MatLab program WriFish.m with 10 populations @ of N = 100 or 1,000 or 10,000 individuals, over 1,000 generations. Compare the visual variance, as above. Note any loss or fixation events.
Allele frequencies continue to vary at random so long as 0 < q < 1: q = 0.0 or 1.0 are absorbing barriers, which once reached end further change. Fate of q in a population is not predictable: between g = 70 & 100 with N = 100, dark orange q goes from 0.70 to 0.15, and light green q from 0.10 to 0.80
HOMEWORK: Repeat the simulations with the MatLab program WriFish.m with 10 populations @ of N = 100 or 1,000 or 10,000 individuals, over 1,000 generations. Compare the visual variance, as above. Note any loss or fixation events.